kffmct
Perform Monte-Carlo testing of a filter used in conjunction with kff.
This makes Monte-Carlo testing very easy to do. It requires that the
filter be set up using the kffoptions structure.
Interfaces
The simplest interface is:
[...] = kffmct(nr, ts, dt, x_hat_0, P_0, options);where nr is the number of simulations to run, ts contains the state
and stop times [t_start, t_stop], dt is the time step, x_hat_0 is
the initial state estimate, and P_0 is the initial covariance.
The true initial state of each run will be randomly displaced from
x_hat_0 by a Gaussian draw from P_0. The simulation will then
propagate and create noisy measurements using the propagation and
measurement functions from the options structure, along with the
appropriate covariance matrices.
If the filter uses an input vector, one can provide a function, u_fcn,
to create an input vector on each sample:
[...] = kffmct(nr, ts, dt, x_hat_0, P_0, u_fcn, options);This function will be called on each time step to produce the input vector used for that step.
Consider covariance can be used too.
[...] = kffmct(nr, ts, dt, x_hat_0, P_0, P_xc_0, u_fcn, options);where P_xc_0 is the initial covariance between the state and consider
parameters.
Making the Monte-Carlo test more interesting is the ability to use
different functions for the truth than for the filter. This allows one
to use a more complex/realistic simulation than the filter uses to
determine how well the filter will function in its real-world
application. To do this, pass the true functions to kffmct.
[...] = kffmct(nr, ts, dt, x_hat_0, P_0, f, Q, h, R, options)where f is the propagation function, Q is the process noise
covariance matrix (or a function that returns the covariance matrix), h
is the observation function, and R is the measurement noise covariance
matrix (or function return such).
The truth can also include u_fcn:
[...] = kffmct(nr, ts, dt, x_hat_0, P_0, u_fcn, f, Q, h, R, options)The different truth can also use different consider covariance.
[...] = kffmct(nr, ts, dt, x_hat_0, P_0, P_xc_0, u_fcn, ...
f, F_c, Q, h, H_c, R, P_cc, options);where F_c and H_c are the Jacobians of the propagation and
observation functions wrt the consider parameters and P_cc is the
consider covariance matrix.
All invocations output the same set of data:
[t, x, x_hat, P, P_xc, y, S, z_hat] = kffmct(...);See the table below for the definition of these outputs.
Here is a summary of the interfaces (dropping subscripts for legibility):
kffmct(nr, ts, dt, xh0, P0, opt);
kffmct(nr, ts, dt, xh0, P0, ufcn, opt);
kffmct(nr, ts, dt, xh0, P0, Pxc0, ufcn, opt);
kffmct(nr, ts, dt, xh0, P0, f, Q, h, R, opt);
kffmct(nr, ts, dt, xh0, P0, ufcn, f, Q, h, R, opt);
kffmct(nr, ts, dt, xh0, P0, Pxc0, ufcn, f, Q, h, R, opt);
kffmct(nr, ts, dt, xh0, P0, Pxc0, ufcn, f, Q, h, R, Pc, opt);
kffmct(nr, ts, dt, xh0, P0, Pxc0, f, Fc, Q, h, Hc, R, Pc, opt);
kffmct(nr, ts, dt, xh0, P0, Pxc0, ufcn, f, Fc, Q, h, Hc, R, Pc, opt);This function by default produces several plots of the results, including the states, measurements, and statistical properties of the errors.
Inputs
nr | Number of runs to make |
|---|---|
ts | Start and stop times, |
dt | Time step |
x_0 | Initial state, either |
x_hat_0 | Initial estimate, either |
P_0 | Initial covariance, |
P_xc_0 | Initial state-consider parameter covariance, |
u_fcn | A function to determine the input vector, |
f | Propagation function, same as for |
F_c_km1_fcn | Jacobian of the propagation function wrt the consider
parameters or a function, as for |
Q_km1_fcn | Process noise covariance matrix or a function, as for
|
h | Observation function, as for |
H_c_k_fcn | Jacobian of the observation function wrt the consider
parameters or a function , as for |
R_k_fcn | Measurement noise covariance matrix or a function, as for
|
P_cc | Consider covariance |
options | The |
(etc.) | Option-value pairs (see below) |
Outputs
x | True states for all runs ( |
|---|---|
x_hat | Estimated states for all runs ( |
P | Covariance matrix for all runs ( |
P_xc | State-consider parameter covariance for all runs
( |
y | Innovation vector for all runs ( |
S | Innovation covariance for all runs ( |
z_hat | Filtered measurements for all runs, formed by passing the
updated estimate to the observation function, producing the
a posteriori expected value of the observation ( |
Option-Value Pairs
'UserVars' | Any additional inputs to pass to any of the user's
functions, such as |
|---|---|
'Plots' | An array indicating which plots should be created automatically use: Use See the engine's Monte-Carlo documentation at http://www.anuncommonlab.com/doc/starkf/engine_tests.html for information on the generated plots. |
'X0' | Initial expected value of true state, which can be
|
'MoveX0' | If the value passed in for |
'Seed0' | By default, Note that you can use this to recreate any specific
run from |
'ConsiderFcns' | By default, where |
Example of a Linear System
Let's use the kff to filter a perfectly linear system.
Define a system.
dt = 0.1; % Time step
F_km1 = expm([0 1; -1 0]*dt); % State transition matrix
H_k = [1 0]; % Observation matrix
Q_km1 = 0.5^2 * [0.5*dt^2; dt]*[0.5*dt^2 dt]; % Process noise covariance
R_k = 0.1^2; % Meas. noise covarianceSet some simulation options.
% Set the initial conditions.
x_hat_0 = [1; 0]; % Initial estimate
P_0 = diag([0.5 1].^2); % Initial estimate covariance
% Define things for kff.
f = @(t0, tf, x, u) F_km1 * x;
h = @(t, x, u) x(1);
options = kffoptions('F_km1_fcn', F_km1, ...
'Q_km1_fcn', Q_km1, ...
'H_k_fcn', H_k, ...
'R_k_fcn', R_k, ...
'f', f, ...
'h', h, ...
'joseph_form', false);Run 20 simulations with different random draws, and then show statistics.
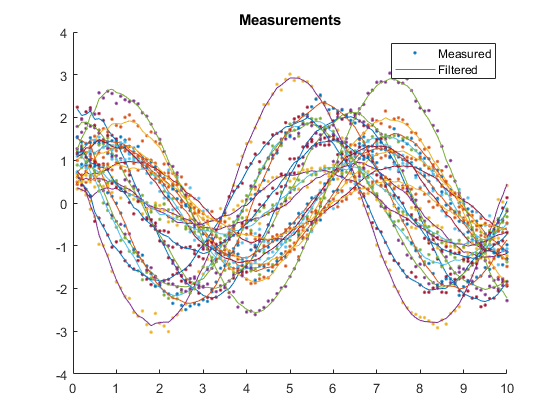
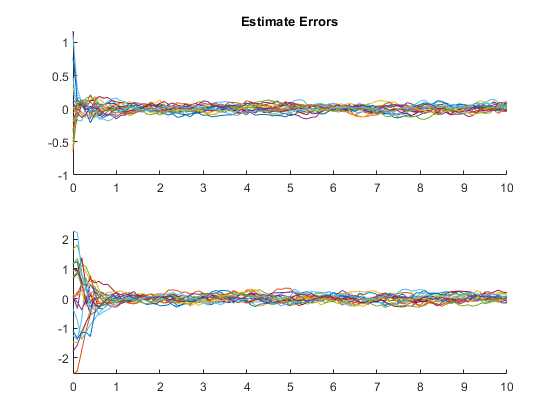
kffmct(20, [0 10], dt, x_hat_0, P_0, options);NEES: Percent of data in theoretical 95.0% bounds: 94.1%
NMEE: Percent of data in theoretical 95.0% bounds: 98.0%
NIS: Percent of data in theoretical 95.0% bounds: 91.0%
TAC: Percent of data in theoretical 95.0% bounds: 94.9%
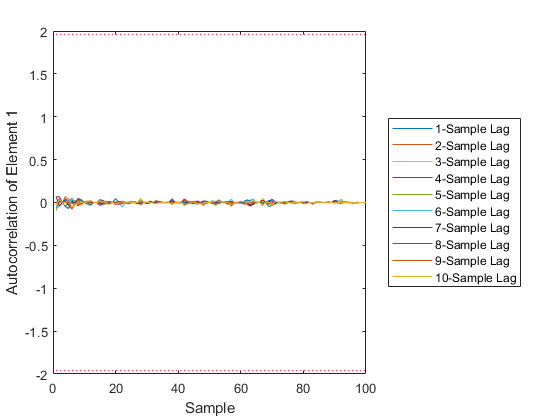
AC: Percent of data in theoretical 95.0% bounds: 95.8%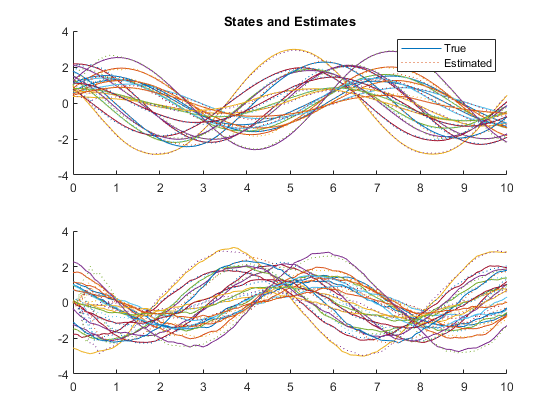
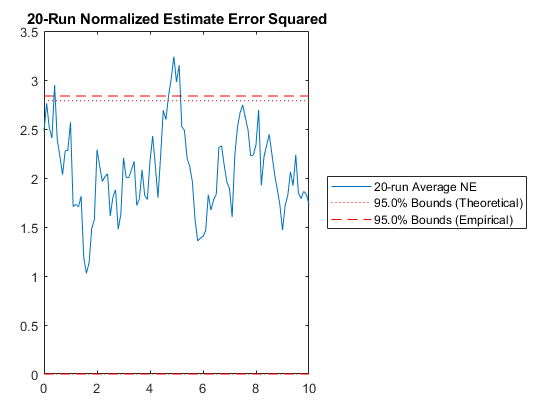
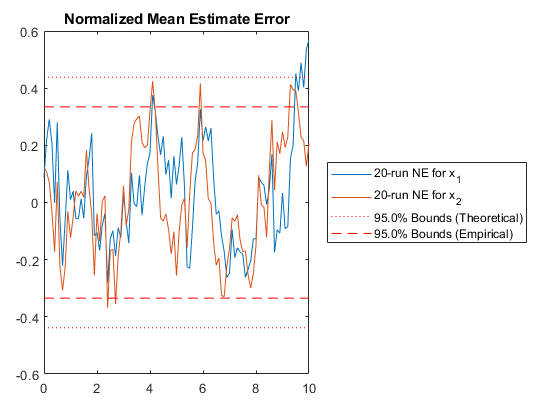
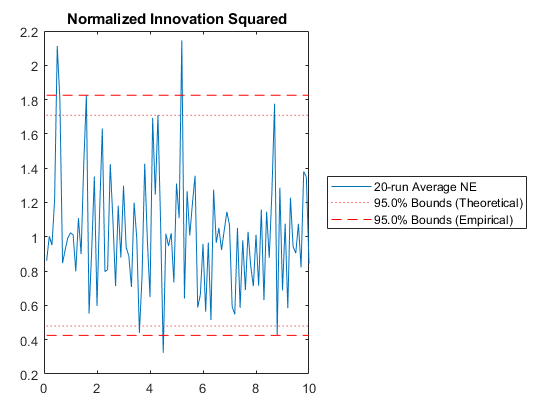
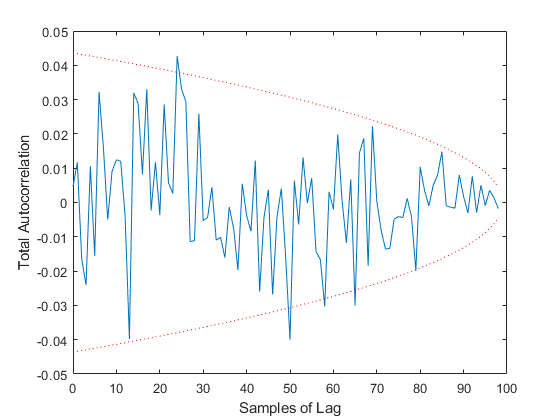
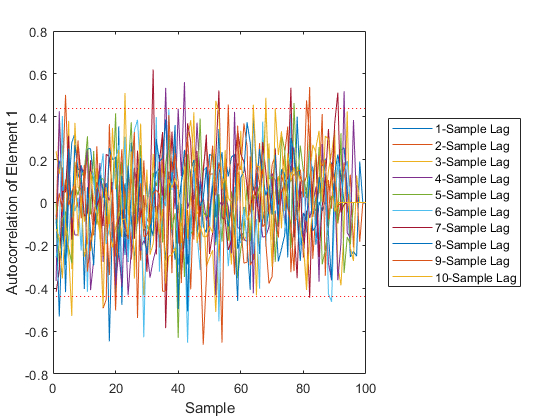
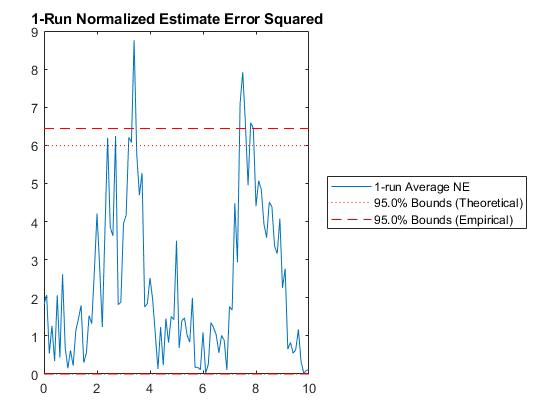
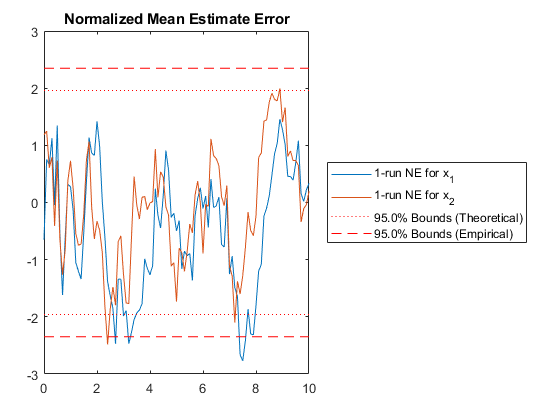
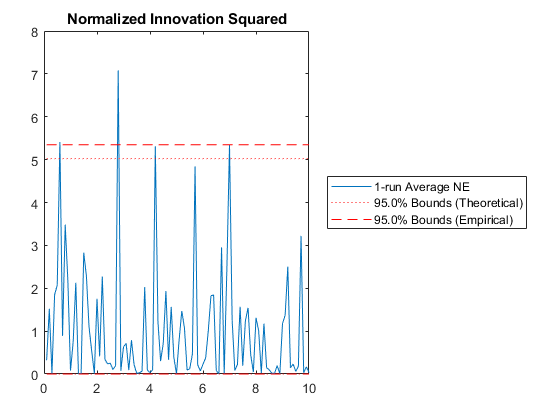
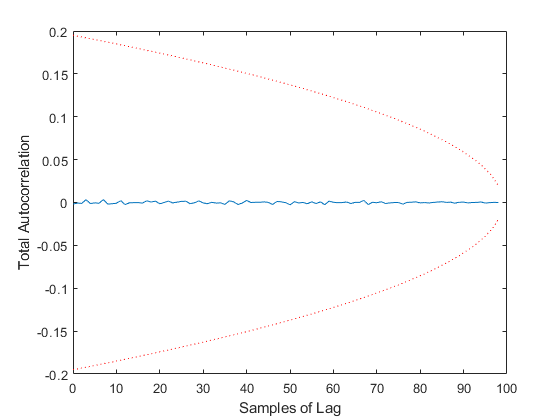
Of course, we can use kffmct to simulate a single run too.
kffmct(1, [0 10], dt, x_hat_0, P_0, options);NEES: Percent of data in theoretical 95.0% bounds: 90.1%
NMEE: Percent of data in theoretical 95.0% bounds: 93.6%
NIS: Percent of data in theoretical 95.0% bounds: 95.0%
TAC: Percent of data in theoretical 95.0% bounds: 100.0%
AC: Percent of data in theoretical 95.0% bounds: 100.0%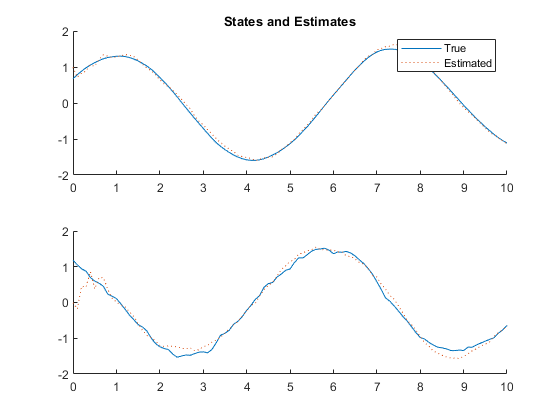
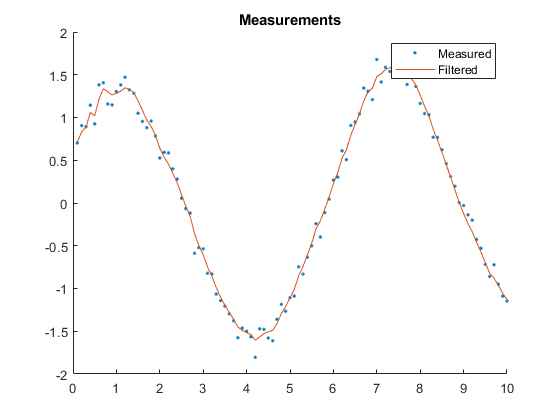
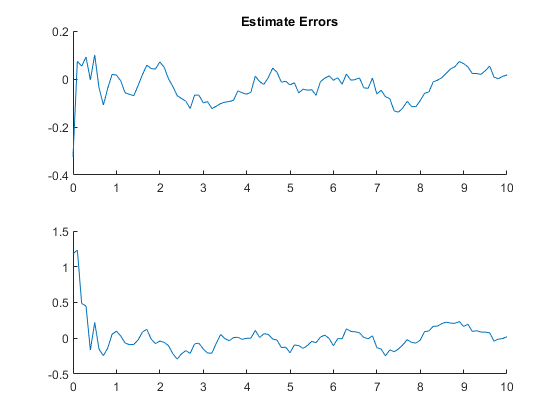
Example of Misalignment between True and Filter Models
We can also use a different model for the true f and h and see what
the result is on the estimator.
f_true = @(t0, tf, x, u) expm([0 1; -0.5 -0.5] * dt) * x; % Different!
h_true = @(t, x, u) 1.5 * x(1); % Also different
Q_true = Q_km1;
R_true = R_k;
% Run 20 sims to see how things look.
kffmct(20, [0 10], dt, x_hat_0, P_0, [], ...
f_true, Q_true, h_true, R_true, options);NEES: Percent of data in theoretical 95.0% bounds: 1.0%
NMEE: Percent of data in theoretical 95.0% bounds: 23.3%
NIS: Percent of data in theoretical 95.0% bounds: 41.0%
TAC: Percent of data in theoretical 95.0% bounds: 28.3%
AC: Percent of data in theoretical 95.0% bounds: 42.0%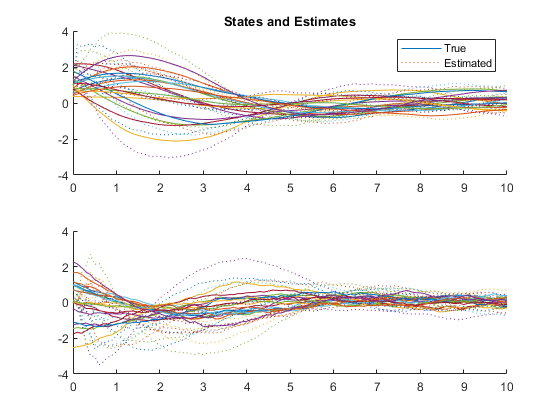
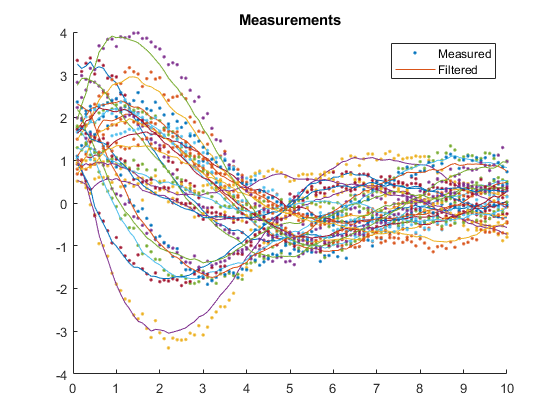
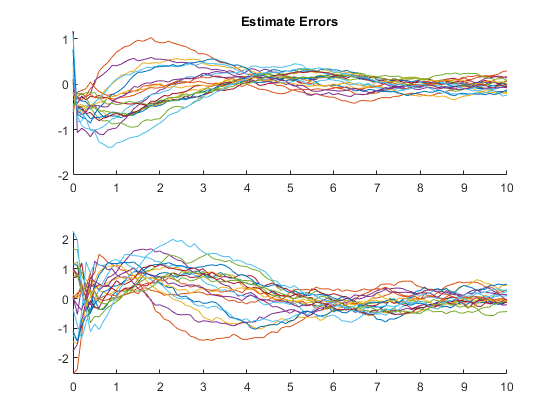
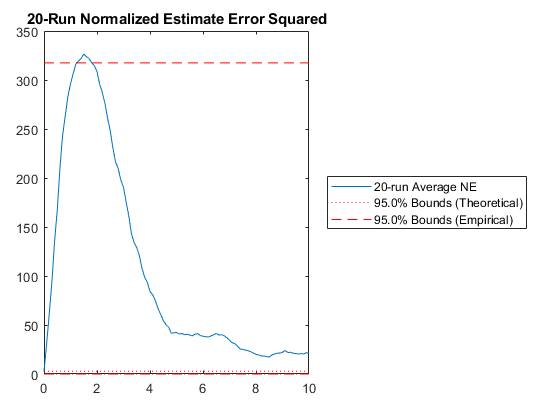
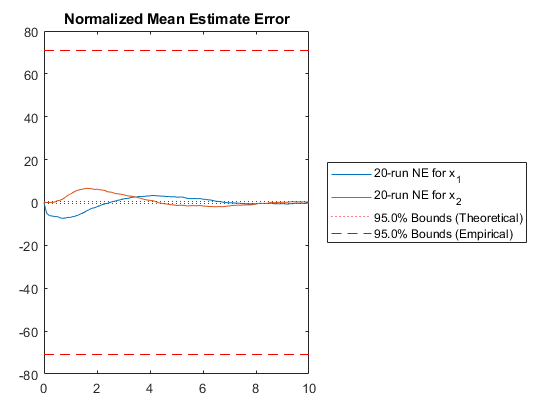
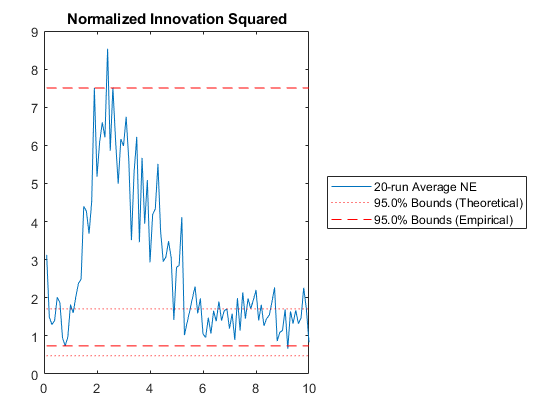
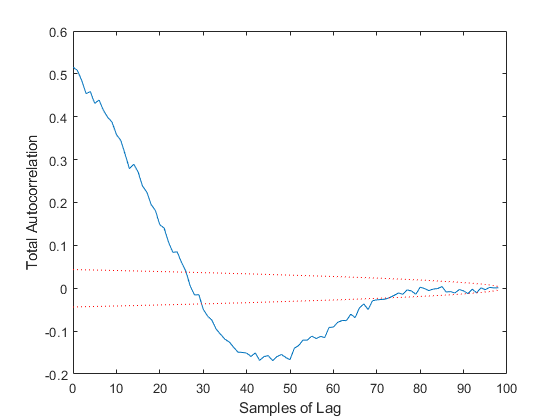
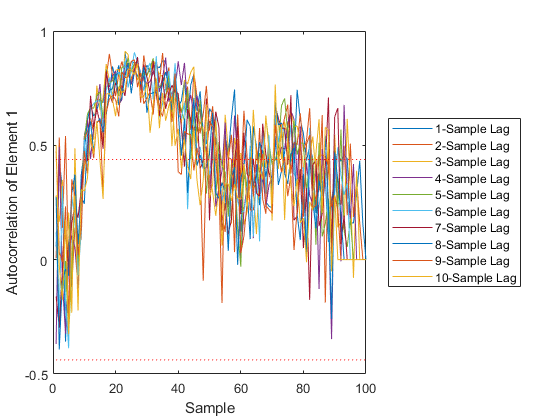
Few points are within the bounds predicted by the estimate covariance, suggesting that the filter is missing some important part of the true system and that our filter is, as a result, too optimistic. We could either figure out more about the true system, where that's feasible, or using consider covariance to represent additional uncertainty in the model.
See Also
*kf v1.0.3 January 17th, 2025
©2025 An Uncommon Lab